clarkia_hz|>
group_by(site)|>
summarise(var_admix =
var(admix_proportion))• 17. ANOVA assumptions
Motivating Scenario: You are ready to run an ANOVA to compare means among more than two groups. Before doing so, you need to check whether your data meet ANOVA’s assumptions (and know what to do if they do not).
Learning Goals: By the end of this subchapter, you should be able to:
List the assumptions of an ANOVA.
Evaluate if your data meet these assumptions.
Identify options for dealing with violations of these assumptions, and know how to choose among them.
Like the two-sample t-test (and all linear models), ANOVA assumes:
- Independence
- Random sampling (no bias)
- Equal variance among groups (homoscedasticity)
- Normality of residuals. As in the two-sample t-test, this requires approximate normality within groups.
Here we use the parviflora admixture data to look into these assumptions
ANOVA assumes…
ANOVA assumes equal variance among groups
Homoscedasticity means that the variance is similar across groups. This makes sense, as we use a mean squares error in ANOVA. In fact, the very derivation of ANOVA assumes that all groups have equal variance.
| site | var_admix |
|---|---|
| SR | 0.01 x 10^4 |
| S22 | 4.02 x 10^4 |
| SM | 0.51 x 10^4 |
| S6 | 1.64 x 10^4 |
As we can see, the variance among groups differs quite substantially. So, we should think twice before conducting a standard ANOVA. Below I will introduce Welch’s ANOVA, to accommodate this issue. But I will nonetheless analyze the data by a standard ANOVA as well to show you the mechanics. At the end we will compare and contrast results from a standard and Welch’s ANOVA to see how worried we should be about equal variance.
ANOVA assumes normal residuals within groups
As in the t-test, and in all linear models, we use the normal framework to evaluate the null hypothesis proposed by an ANOVA. Of course data are rarely perfectly normal, but the data we are examining here seem particularly far from this assumption (see Figure 1). I will proceed here, not because this is the right thing to do, but simply to show you how an ANOVA works.
Code
library(patchwork)
qq_clarkia_hz <- clarkia_hz|>
ggplot(aes(sample = admix_proportion))+
geom_qq()+
geom_qq_line()+
facet_wrap(~site, scales = "free_y", nrow = 1)
hist_clarkia_hz <- clarkia_hz|>
ggplot(aes(x = admix_proportion))+
geom_histogram( bins = 10, color = "white")+
facet_wrap(~site, nrow = 1, scales = "free_y")
qq_clarkia_hz / hist_clarkia_hz + plot_annotation( tag_levels = "A" )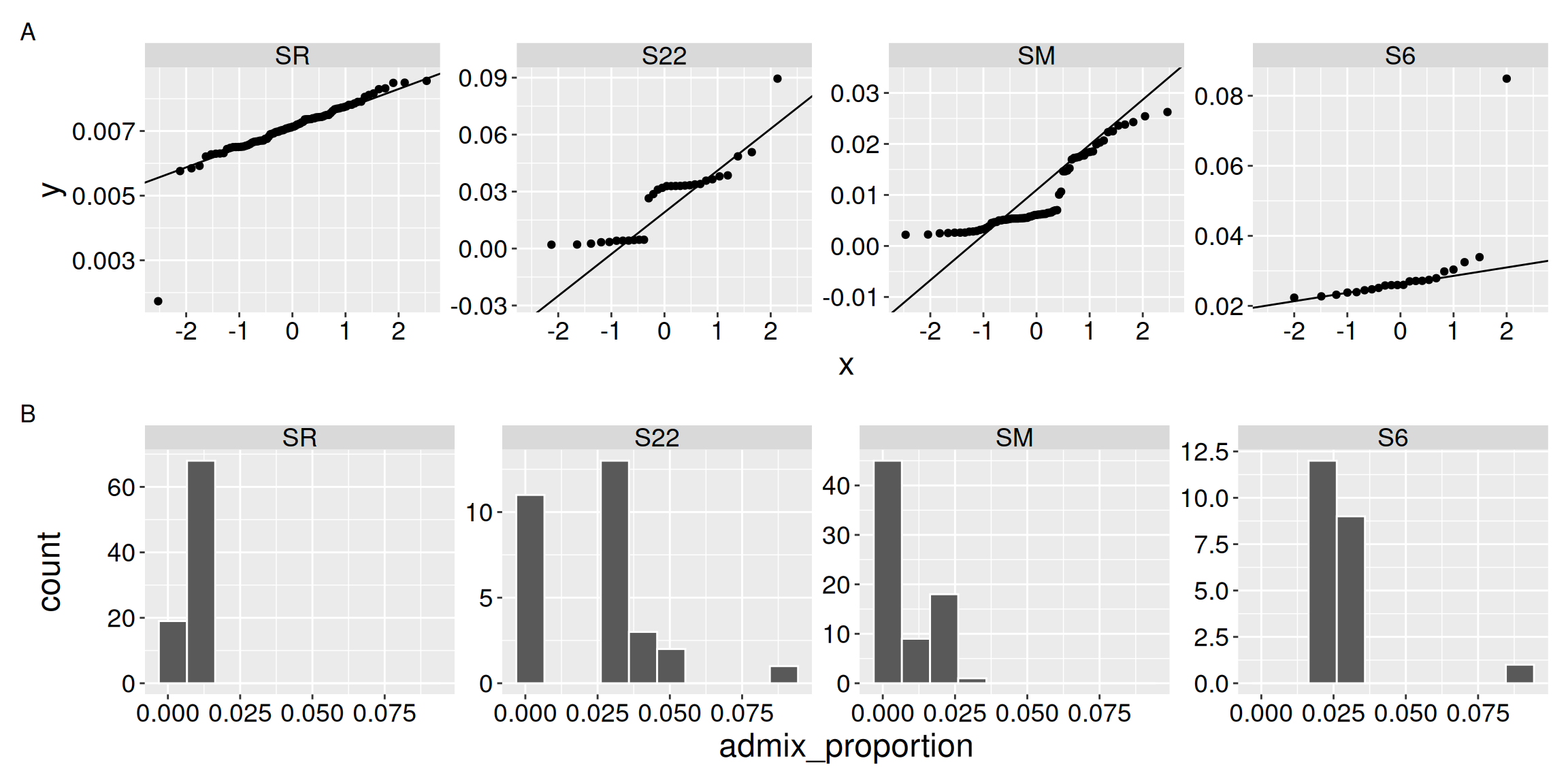
ANOVA assumes independence within groups
Ugh the news gets even worse. These samples are taken from a reasonable proportion of the extant plants and have a spatial structure to them (Figure 2). My sense is that they are siblings or even genetically identical “clones”. The appropriate statistical procedure is not clear, but we will hold our nose, do our best, and acknowledge the potential non-independence.
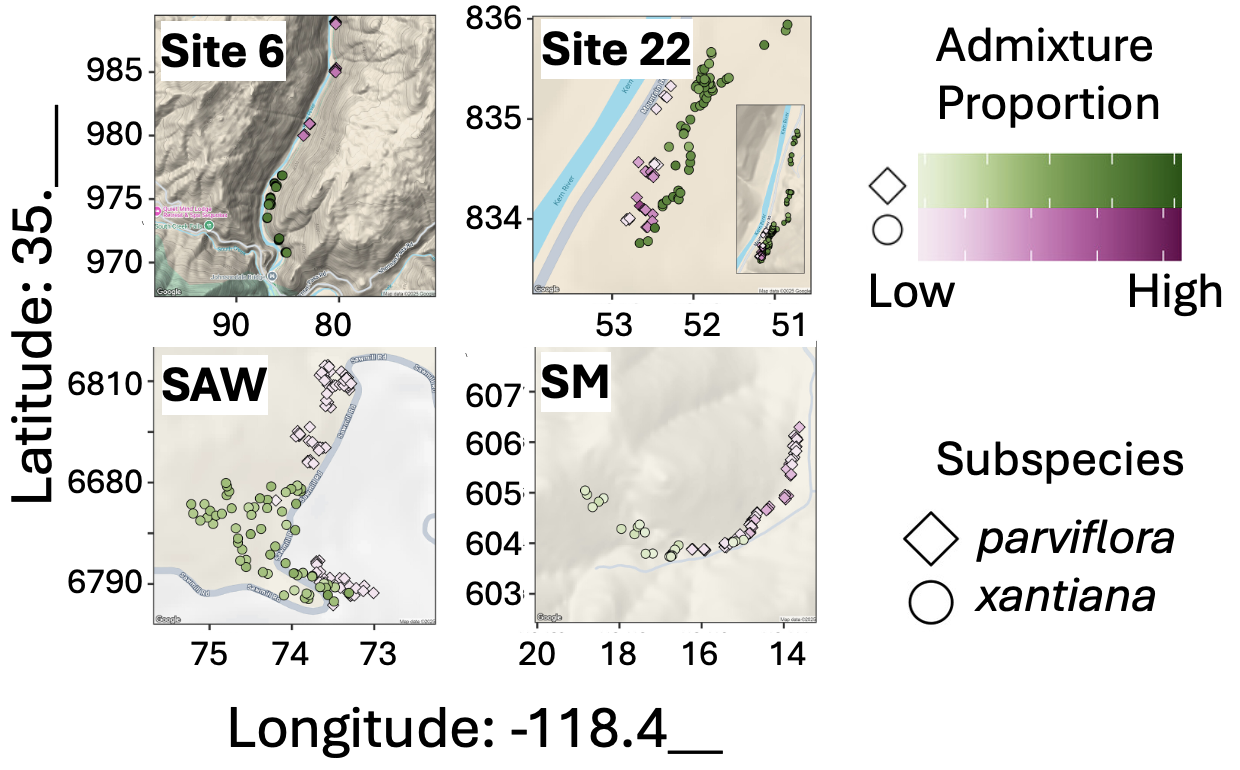
ANOVA assumes unbiased sampling within groups
We did an awesome job here. Don’t worry about this one!
What to do when you violate assumptions
We’ll continue using these data to learn how ANOVA works. But when the assumptions are clearly broken, as in this example, we need to think about what to do. I want to emphasize that there’s usually not a single right answer, and that your choice will be a compromise between trade-offs of each potential approach. Below are some options, along with my (strong) opinions. I also note that you don’t need to pick a single analysis, sometimes it makes sense to present results from different types of analyses to show that your take-home message is robust to any specific assumptions.
Most importantly, think about what you’re doing and why. While you should not run a statistical test that gives meaningless results, there’s usually a trade-off between statistical purity and clear communication of your results. Don’t rush to the most statistically pure approach, if it means your results are difficult to interpret.
If you’re interested in NHST, breaking assumptions means that your p-values aren’t properly calibrated, so you cannot literally interpret your p-value. But how often do you care about the exact p-value? If you’re worried about misleading confidence intervals, consider the trade-offs between having these perfectly calibrated, and loss of information that may occur when presenting results on a scale that is difficult to interpret. Let clarity about your goals and the meaning of your results take precedence over strict rule following.
Minor violations: If the test is known to be robust to small deviations—like mild non-normality or modest differences in variance—you can usually proceed without concern.
Major non-normality: When data are far from normal within groups, consider transforming your data (e.g., log, square root) or using a model that explicitly captures the shape of the distribution that generated your data. We’ll return to this idea later when we discuss generalized linear models.
Robust approaches: Methods like Welch’s ANOVA (which allows group variances to differ, below), or approaches that “trim” extreme outliers, are generally solid and defensible choices.
Permutation tests: are often appealing because they avoid assumptions about the distribution of the data. But they still require independence within groups. A simple permutation test cannot fix this problem. As always, no method gets you out of thinking carefully about how your data were generated.
Rank-based tests: Alternatives like the Kruskal–Wallis test assess differences in medians rather than means and are less sensitive to assumption violations. However, their effect sizes are harder to interpret in practical or biological terms.
Welch’s ANOVA when variance differs
You can use Welch’s ANOVA when variance differs substantially among groups. Welch’s ANOVA has the same underlying logic as a standard ANOVA, but relaxes the assumption of equal variance. Specifically, it adjusts the calculation of the test statistic and degrees of freedom to account for differences in group variances, effectively giving less influence to groups with higher variance. This means that F and the degrees of freedom etc can be interpreted similarly as in a standard ANOVA, but are calculated differently.
The oneway.test() function in R uses Welch’s ANOVA by default:
oneway.test(admix_proportion ~ site, data = clarkia_hz)
One-way analysis of means (not assuming equal variances)
data: admix_proportion and site
F = 32.224, num df = 3.00, denom df = 52.34, p-value = 6.117e-12Here, we resoundingly reject the null hypothesis that all groups have the same mean.
